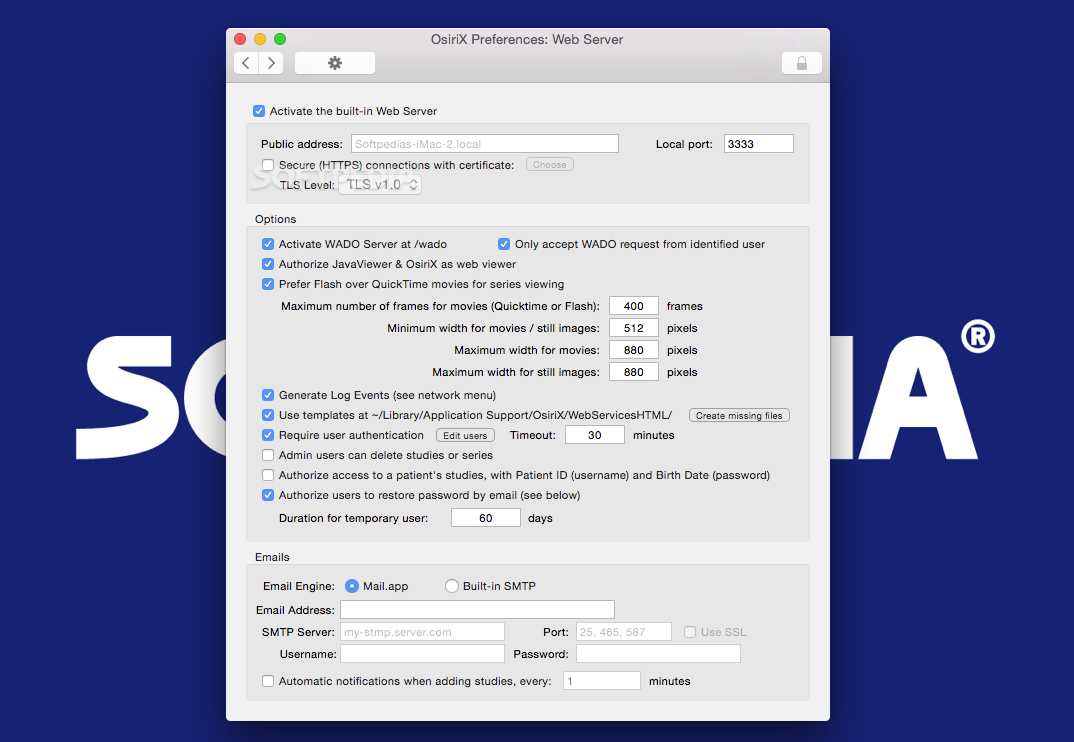 
These image labels are human-readable, but not machine-accessible. This image is annotated to convey the major semantic content, including anatomy (right lobe of liver), pathology (irregular mass, tumor vascularity), and imaging observations (2 cm in size, irregular shape, and poorly-defined margins). Radiological image with semantic annotations drawn in an image annotation tool. A tool permitting researchers to describe the semantic information in images in a manner that fits within the research workflow could help them to collect the necessary structured image data. While these systems provide tools to label images ( Figure 1), they do not permit users to record the key semantic image content in a standard machine-interpretable format. Images are viewed and interpreted in Picture Archiving and Communication Systems (PACS) or in a variety of image viewers for image manipulation. 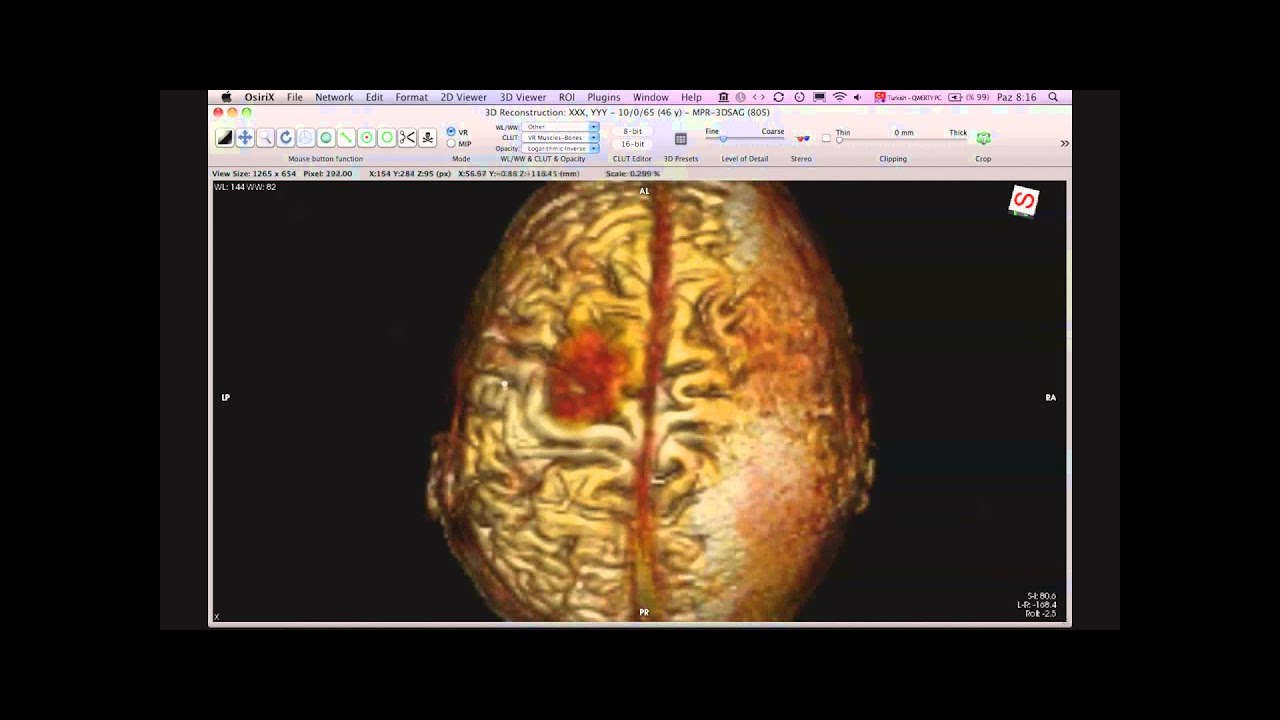
Schemas to make the semantic contents of images explicit are being developed, 1 – 3 however, experts who interpret images do not generally record their observations in the highly structured format of such schemas. In addition, it would be possible to combine images with complementary non-image data to enhance biomedical discovery. If the semantic information in images were recorded in a structured manner, researchers and clinical practitioners could more efficiently search and analyze large image repositories according to particular features of images (e.g., discover characteristic image appearances of particular diseases, compute the average lesion size, and summarize types of abnormalities in different diseases). Thus, much critical radiological image information is not directly accessible to machines, unlike other biomedical data such as genomics, proteomics, and electronic medical records data. 
The radiologist-extracted semantic information is rarely linked or embedded within the images, instead being recorded in text reports or in case report forms as part of clinical trials. The “semantic content” in images is the information extracted by radiologists who view them: the visual appearance of anatomic structures and abnormalities contained in those structures (“image findings” such as a mass in the liver or an enlarged heart). While the raw images themselves can be useful in some computer processing applications, much of the critical semantic content in images is not explicit and available to computers, hindering discovery in large-volume image data sets. Images produced by radiology modalities are a critical data type in biomedicine because they provide rich visual information about disease, complementary to non-image data such as genomics, proteomics, histopathology, and clinical laboratory results. Tools such as iPad can help reduce the burden of collecting structured information from images, and it could ultimately enable researchers and physicians to exploit images on a very large scale and glean the biological and physiological significance of image content. Image annotations are saved in a variety of formats, enabling interoperability among medical records systems, image archives in hospitals, and the Semantic Web. iPad hides the complexity of the underlying image annotation information model from users, permitting them to describe images and image regions using a graphical interface that maps their descriptions to structured ontologies semi-automatically. We have created iPad, an open source tool enabling researchers and clinicians to create semantic annotations on radiological images. Information schemes are being developed to describe the semantic content of images, but such schemes can be unwieldy to operationalize because there are few tools to enable users to capture structured information easily as part of the routine research workflow. Radiological images contain a wealth of information, such as anatomy and pathology, which is often not explicit and computationally accessible.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
